This webpage was produced as an assignment for Genetics 677, an undergraduate course at the University of Wisconsin - Madison.
Homologous Protein Alignments
ClustalW
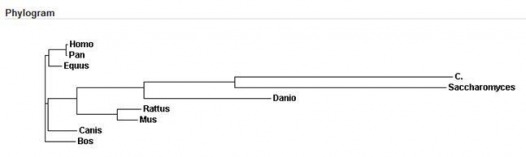
ClustalW is a general purpose multiple sequence alignment program for DNA or proteins. It produces biologically meaningful multiple sequence alignments of divergent sequences. It calculates the best match for the selected sequences, and lines them up so that the identities, similarities and differences can be seen. Evolutionary relationships can be seen via viewing cladograms or phylograms. (1)
The following file is the ClustalW alignment of the Homo sapiens CFTR protein to the homologous gene in Pan troglodytes, Equus caballus, Canis lupus familiaris, Bos taurus, Rattus norvegicus, Mus musculus, Danio rerio, Saccharomyces cerevisiae, and C. elegans. All default settings were used.
| clustalw_protein_alignment.txt |
MUSCLE
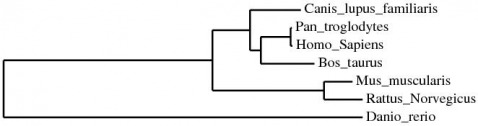
MUSCLE stands for MUltiple Sequence Comparison by Log-Expectation. MUSCLE is claimed to achieve both better average accuracy and better speed than ClustalW2 or T-Coffee, other protein alignment programs.
The following file is the MUSCLE alignment of the Homo sapiens CFTR protein to the homologous gene in Pan troglodytes, Equus caballus, Canis lupus familiaris, Bos taurus, Rattus norvegicus, Mus musculus, Danio rerio, Saccharomyces cerevisiae, and C. elegans. All default settings were used.
| muscle_protein_alignments.doc |
Analysis
All alignments matched the sequences very similarly, their divergence appropriate to their evolutionary development. The regions of greatest dissimilarity between the homologs were the C terminal and N terminal.
Rattus norvegicus and Saccharomyces cerevisiae both have extended N-terminal sequences of at least 55 amino acids. Saccharomyces cerevisiae and C. elegans both have shortened C-terminal sequences and have the most gaps compared to any other homolog; which is expected since they are so divergent from mammals.
Rattus norvegicus and Saccharomyces cerevisiae both have extended N-terminal sequences of at least 55 amino acids. Saccharomyces cerevisiae and C. elegans both have shortened C-terminal sequences and have the most gaps compared to any other homolog; which is expected since they are so divergent from mammals.
_______________
References
1. ClustalW. Accessed March 18, 2010. http://www.ebi.ac.uk/Tools/clustalw2/index.html
2. MUSCLE. Accessed March 19, 2010. http://www.ebi.ac.uk/Tools/muscle/index.html
References
1. ClustalW. Accessed March 18, 2010. http://www.ebi.ac.uk/Tools/clustalw2/index.html
2. MUSCLE. Accessed March 19, 2010. http://www.ebi.ac.uk/Tools/muscle/index.html
Alexandra Reynolds
[email protected]